Rule-based modeling allows for the construction and simulation of models representing chemical interactions within and between cells. However, rule-based modeling has several inherent disadvantages. For one, systems biologists are not software developers – spotting syntax and semantic errors in the model file may be extraordinarily difficult. Additionally, the numerical output of model simulations may be difficult to interpret.
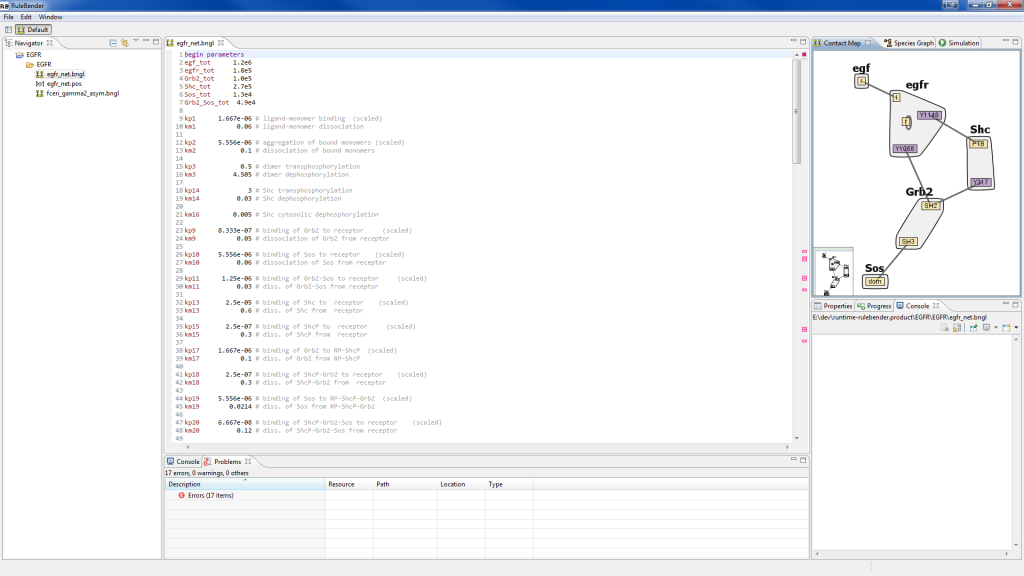
RuleBender is essentially a development environment for the BioNetGen language, containing a visual debugger, model simulator, and results viewer. The goal of this project is to facilitate rule-based modeling construction, simulation, and analysis in an integrated system.
More information about the project, as well as a download link, can be found at the official RuleBender website.